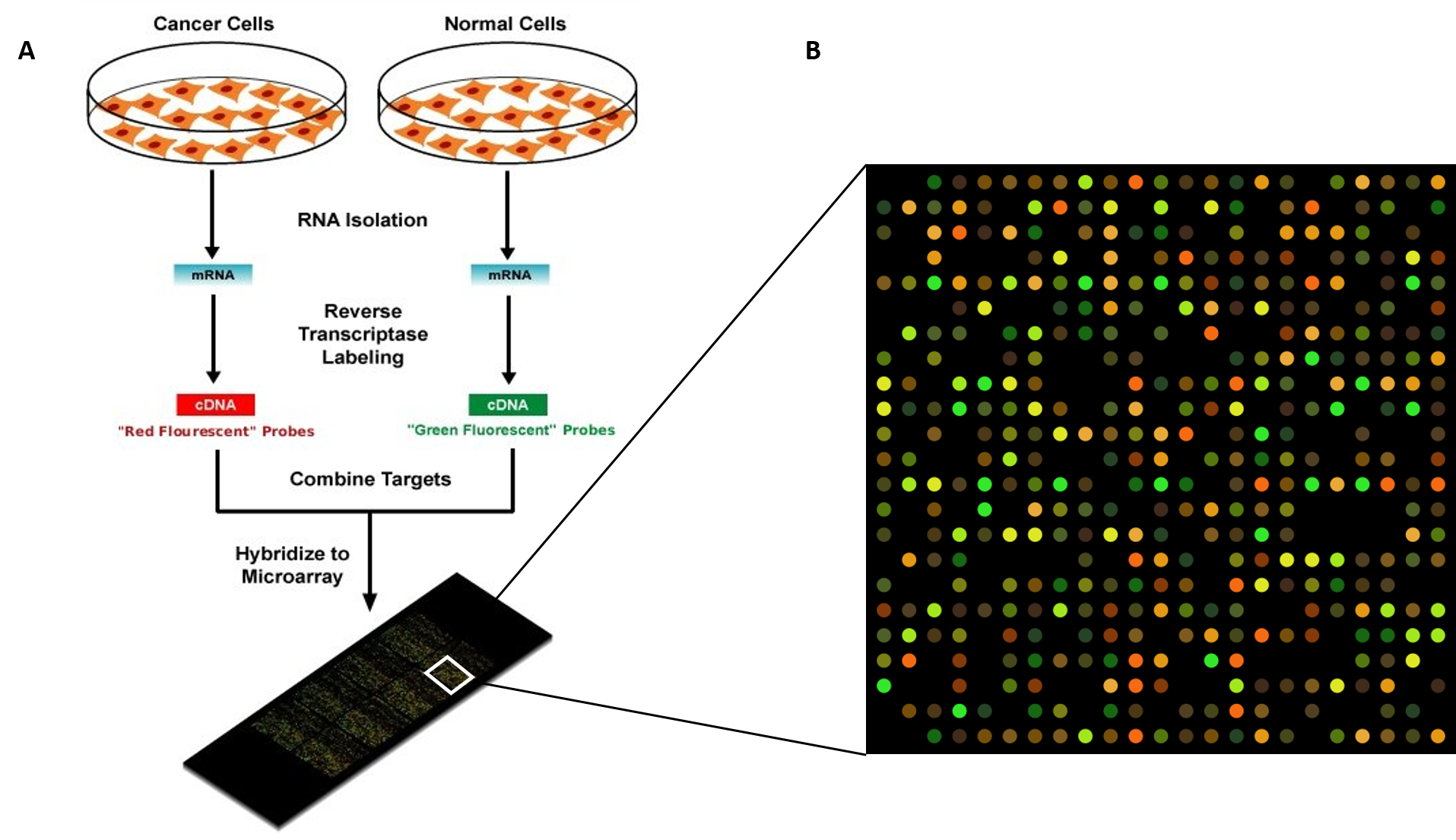
How cool! Brain clusters with brain, and liver with liver. Heatmap it up! heatmap(exprSet.nologs, main = "Normalized ME matrix for brain, liver, N=8") Now we have normalized expression values, and we can look at the expression matrix! That was so darn easy. 
We then correct for negative values, and the signals are scaled based on this mean, meaning that we have background adjusted values. We then take the mean of the weighted intensity values and that becomes the mean for our Kth set. The probe pair gets weighted based on the difference from the mean. Using the two, meaning that if MM > PM, we get negative values. We calculate an average difference intensity value The standard deviation of the lowest 2% cell intensities is calculated to estimate variabilityĪfter much digging, I finally found good documentation for PM/MM - it's referring to a probe set that includes a Perfect Match (PM) and MisMatch to a reference sequence. Rank the cells, the loweset 2% are chosen as the background b for the zone.ģ. splitting the array into K (default K=16) rectangular zones, not including control or masked cells.Ģ. Specifically, we calculate a background “floor” to subtract from each cell value. This is what's going on, accoring to the script Summarize and normalize with MAS5 eset.mas5 = mas5(affy.data) CEL files files = list.files("/home/vanessa/Documents/Work/GENE_EXPRESSION/tutorial", 
# 'citation("Biobase")', and for packages 'citation("pkgname")'. # Vignettes contain introductory material view with # rbind, Reduce, rep.int, rownames, sapply, setdiff, sort, # order, paste, pmax, pmax.int, pmin, pmin.int, Position, rank, # intersect, is.unsorted, lapply, Map, mapply, match, mget, # colnames, duplicated, eval, evalq, Filter, Find, get, # anyDuplicated, append, as.ame, as.vector, cbind, # The following objects are masked from 'package:base': # The following object is masked from 'package:stats': # parLapplyLB, parRapply, parSapply, parSapplyLB # clusterExport, clusterMap, parApply, parCapply, parLapply, # clusterApply, clusterApplyLB, clusterCall, clusterEvalQ, # The following objects are masked from 'package:parallel': Set your working directory to where we downloaded tthe files setwd("/home/vanessa/Documents/Work/GENE_EXPRESSION/tutorial") # Old packages: 'httpuv', 'IRanges', 'oligo' # Warning: installed directory not writable, cannot update packages 'boot', # Using Bioconductor version 2.13 (BiocInstaller 1.12.0), R version 3.0.2. # Bioconductor version 2.13 (BiocInstaller 1.12.0), ?biocLite for help try to find what functions specific groups of genes (with similar expression profiles) have in commonįirst, fire up R and bioconductor source("").cluster a differentially expressed subset of all genes to identify those with similar expression profiles.flag low intensity data (most probably background noise).use a common statistical test to identify differentially expressed genes.calculate expression ratios of genes between two different tissues.calculate Absent/Present calls which attempts to label genes that are "expressed”.preprocess the raw data (summarize probe measurements into one measurement for each probe).You can download and zip to a directory of your choice, and fire up R. I am following along directly from that linked page. Let's get the data, from here These arrays were “hybridized to tissues from fetal and human liver and brain tissue, courtesy of MIT. For Affymetrix methods (implemented in Bioconductor), I will reference this document Preparing for Analysis with R I'm not going to claim to know anything about the preparation of the DNA - that's for the wet lab folk :) What is important to me is that this machine can scan the array, analyze the images, and spit out expression profiles For more details, here is the reference I was reading. What in the world? Basically, we can evaluate mRNA expression of thousands of genes at once, which are put on one of those cool chips. The raw data is in the form of “.CEL” files, which I believe are produced by the Affymetrix GeneChip System. I've been working with microarray expression data from the Broad Connectivity Map, and of course decided to download raw data (to work with in R) over using their GUI. This is an affymetrix tutorial, for analysis of.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
